Four Powered Online Tools for Genomic Analysis and Visualization (I) – getorf
I.getorf – ORF prediction
Open Reading Frames (ORFs) represent a region of specified minimum size between two STOP codons, or between a START and a STOP codon. To predict and annotate ORFs in a sequence, ORF prediction tools are commonly used. getorf is an (online) software tool that finds and extracts open reading frames present in any sequence that a user can feed as an input. It is a command line program from EMBOSS (the European Molecular Biology Open Software Suite), and a part of Nucleic: Gene finding command groups.
getorf can work with either a single or multiple nucleotide sequence. The program takes a standard EMBOSS sequence query (also known as ‘USA’) as an input which mainly includes srs:embl, srs:uniprot and esembl, as defined in EMBOSS installations. Alternatively, data can also be read from sequences written by an EMBOSS or other third-party application as long as it is a supported format. The format of the input can be specified using command-line qualifier – sformat ‘xxx’, where ‘xxx’ is replaced by the format name. Formats that are available are: gff (gff3), gff2, embl (em), genbank (gb, refseq), ddbj, refseqp, pir (nbrf), swissprot (swiss, sw), dasgff and debug.
Once the input format and sequence are defined, then the tool can be used in a pretty straight-forward way from the interface, as below. The search can customize by different parameters such as the organism, minimum/maximum size of the ORF to report, the type of sequence (circular/linear), number of flanking nucleotides to report. Additionally, the type of output and its format can also be defined according to the need. When dealing with relatively larger sequences, one can opt to receive the results in email as well[1].
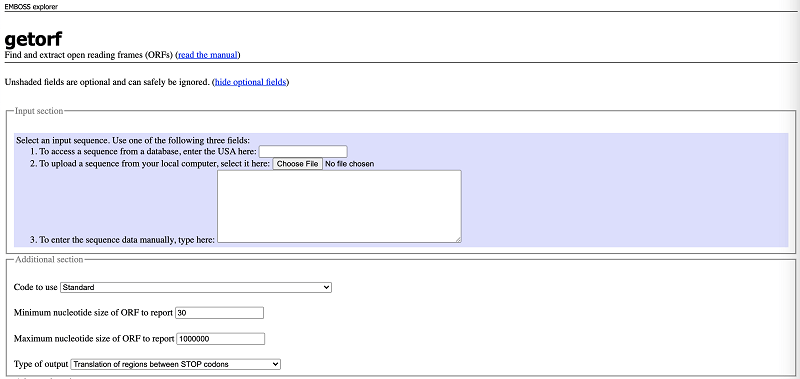
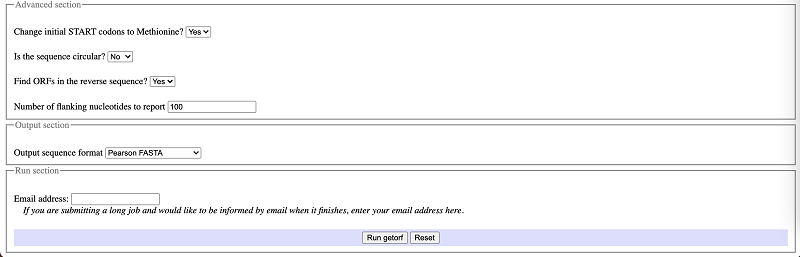
There are three ways to upload the sequences. After sequence input, click ‘Run getorf’ and you will get output like the sequence shown below. And finally copy and save the results and you can use blastp to do the successive analysis.
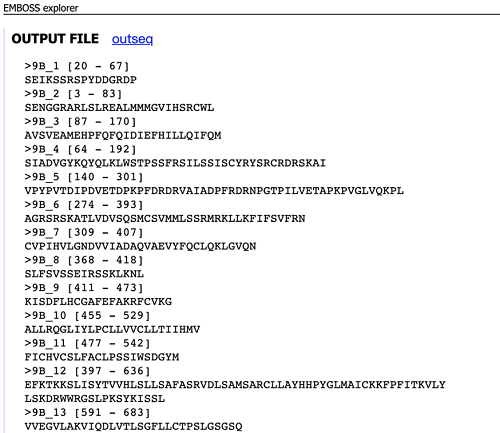
To be continued…
Reference:
[1] http://emboss.sourceforge.net/apps/cvs/emboss/apps/getorf.html
