A Beginner’s Guide to Microbial Shotgun Metagenomic Sequencing
Shotgun metagenomic sequencing uses modern genomic techniques to sequence all the genes in all the microorganisms present in a sample without the need for specific isolation or cultivation of individual species. This opens a world of research as there are between 1030 and 1035 microbial genomes, and nearly 99% of all microorganisms cannot be cultivated in the laboratory. This method enables researchers to evaluate microbial diversity, detect the abundance of microbes in the environment sampled, and investigate microbial structure and interactions. In addition, the sequencing results can also be used to assess functional diversity in the sample analysed.

Shotgun metagenomic sequencing is faster and more reliable than traditional techniques as it removes the need for culturing microbes which could create conditions that favour certain species and introduce bias into the results. Further, metagenomic sequencing removes other sources of bias associated with traditional techniques, including multiple PCR amplification requirements.
Shotgun Metagenomic Sequencing Workflow
A shotgun metagenomics project workflow typically follows the same six steps: sample preparation, DNA extraction, Quantification and Quality control, Library construction, sequencing, and bioinformatics analysis.
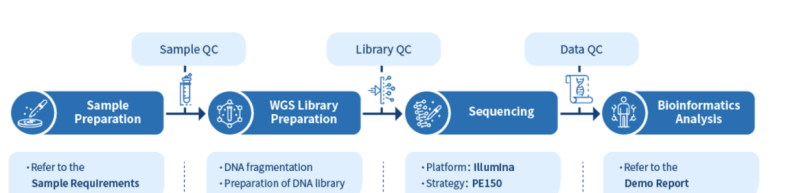
Sample preparation depends on the environment that you are sampling from. The samples generally taken for these types of projects range from soil and water to faeces, swabs, and tissues. Some issues can arise with sample preparation, especially those taken from tissues, as they may have high risk of contamination with the host genome. This can result in low-quality data being produced. Certain strategies can be used to avoid this, such as not sampling too close to the host’s tissue and using the correct kit when extracting. In addition, if the host’s genome is known, it is possible to remove this from the sequencing data.
Once the sample has been prepped, DNA can be extracted. The preferred method for DNA extraction is using CTAB method. However, depending on the specificality of your samples, some customized kits may work better, for example, for sludge and soil samples, it is highly recommended using the PowerSoil ® DNA isolation kit. Following extraction, the samples will then undergo quality control to check whether there is any degradation and the samples are good enough to proceed to library prep.
DNA is fragmented into 250 – 300 bp length for constructing a library of 350bp insert size. This library is then sequenced using the Illumina Novaseq platform with pair-end 150 bp strategy. Once sequenced, the reads then go through data quality control to filter the reads. There are three types of read that are removed:
- Reads that contain adapters
- Reads containing >10% of unknown bases
- Low-quality reads
After data filter, the clean reads can proceed to the analysis pipeline.
With shotgun metagenomic sequencing data, we are interested in answering two types of questions. The first is to examine the sample to see who is there, and the second is to see what they are doing. More specifically, to answer the first question, we are interested in examining the taxonomic and phylogenetic diversity to see what types of microbes are present in the sample. The second question addresses the role the microbes play within the sample by looking at gene prediction and functional annotation. We first need to assemble the reads and annotate the results to answer these questions. Following this, a host of downstream analyses can be carried out depending on the samples and questions you have. These include but are not limited to:
- Gene catalogue statistics
- Taxonomy annotation
- PCA and NMDS Analyses
- Functional Annotation
- Antibiotic-Resistant Gene Distributions
Shotgun Vs Amplicon Sequencing
Both shotgun and amplicon sequencing can provide in-depth information about microbial communities, but which one you use depends on the research questions you are asking. 16S/18S/ITS Amplicon Metagenomic Sequencing focuses on sequencing specific target regions of DNA and can be used to screen variants or target specific organisms. This sequencing method is commonly used in population and community microbial ecology studies to identify individual species within a pure culture and detect organisms of interest, such as pathogens.
In contrast, shotgun metagenomic sequencing provides genetic information on all the organisms in a sample. Using this method, the total DNA from the sample is sequenced using next-generation sequencing (NGS) technology. This approach is especially effective because it avoids the need for the isolation and cultivation of microorganisms.
How Novogene Can Help
Novogene is a world leader in NGS services and has a vast amount of experience providing metagenomic sequencing services. We can offer a range of services from DNA extraction right through to sequencing and bioinformatics, and our team of experts is always on hand to answer any questions you may have. In addition, Novogene has recently introduced Falcon/FalconII, a fully automated, agile, and intelligent delivery system that operates 24/7, reducing both time and costs.
For more information on our shotgun metagenomic sequencing, be sure to have a look at our webinar available here.
Meta description: Shotgun metagenomic sequencing uses modern genomic techniques to sequence all the genes in all the microorganisms present in a sample without the need for specific isolation or cultivation of individual species.
