Illumina Fluent PIP-seq Single Cell 3’ RNA Sequencing
At Novogene, we offer end-to-end Illumina® Fluent PIP-seq single-cell RNA sequencing services, encompassing optimized sample handling, high-quality library construction, NovaSeq™ X Plus sequencing, and comprehensive DRAGEN-accelerated bioinformatic processing within a fully integrated workflow.
This robust single-cell transcriptomic service enables in-depth characterization of cellular heterogeneity, delineation of cell types and states, identification of rare and transitional populations, and exploration of dynamic transcriptional programs across diverse biological contexts, including immunology, development, oncology, neuroscience, and translational research.
Illumina Single Cell 3’ RNA Technology
Illumina Single Cell 3’ RNA Technology employs cell barcoding and UMI-based quantification to efficiently capture poly(A)+ transcripts at single-cell resolution, ensuring precise and reproducible gene expression measurements for each individual cell. Sequencing outputs are processed through the DRAGEN pipeline to produce standardized gene-by-cell expression matrices that are fully compatible with leading downstream analysis platforms such as Seurat and Scanpy.
The Illumina Single Cell platform includes scalable configurations (T2, T10, T20, and T100) to accommodate varying project sizes and throughput requirements. At Novogene, we currently deliver this workflow using the Illumina Fluent™ T10 platform, providing standardized library preparation, scalable throughput, and consistent performance across projects.
To further assess platform performance, Novogene conducted an internal benchmarking study evaluating gene detection sensitivity, UMI distribution, mapping efficiency, and clustering consistency across single-cell platforms. The findings demonstrated strong concordance in transcriptomic resolution and downstream analytical compatibility. Detailed benchmarking results are available in our technical poster.
Why Partner with Novogene for Illumina Single Cell 3' RNA Sequencing?
Deep Technical Experience
Our teams bring extensive hands-on experience in transcriptomic sequencing projects, ensuring robust experimental design, precise execution, and dependable data generation across a broad range of research applications.
Advanced Sample Handling Support
We provide flexible sample preparation options, including nuclei isolation and tailored workflows for frozen specimens. These approaches enable reliable single-cell gene expression profiling from complex or preservation-sensitive samples.
Integrated End-to-End Illumina Workflow
Novogene offers a seamless, end-to-end Illumina-based single-cell service — from sample preparation and library construction to NovaSeq™ X Plus sequencing and DRAGEN-enabled data processing. This unified approach ensures efficient project management, technical consistency, and delivery of standardized, analysis-ready datasets.
Efficient and Practical Approach
Designed as a streamlined single-cell solution, this service balances performance and affordability. Optimized laboratory procedures and scalable sequencing capacity allow researchers to obtain high-resolution cellular insights within practical budget considerations.
Illumina scRNA-seq Specification: Sample Requirements
| Category | Supported Options |
| Species | Human, Mouse, Rat, Other species* |
| Sample Type** | Fresh/Cryopreserved Cell Suspension, Fresh/Frozen Tissue |
*Please contact our team for species-specific handling recommendations and optimal sample preparation guidance.
**Sample compatibility may vary depending on library preparation requirements.
Illumina scRNA-seq Specification: Sequencing and Analysis
| Sequencing Platform | Illumina NovaSeq Xplus | |
| Read Length | Paired End (PE) 150 bp | |
| Data Analysis Capability | DRAGEN Analysis |
|
| Standard Analysis |
|
|
| Advanced Analysis |
|
|
Service Workflow Overview
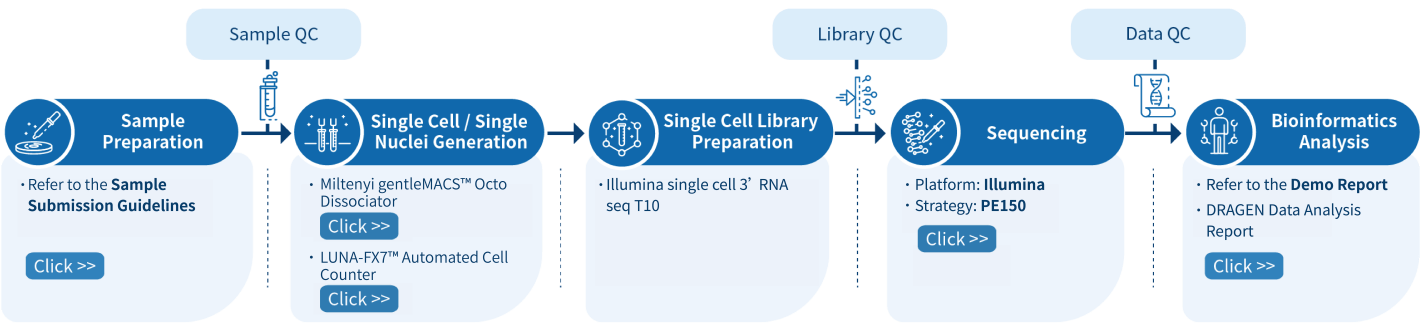
-
Dysregulation of brain and choroid plexus cell types in severe COVID-19
Yang, A.C., Kern, F., Losada, P.M. et al. Dysregulation of brain and choroid plexus cell types in severe COVID-19. Nature 595, 565–571 (2021). https://doi.org/10.1038/s41586-021-03710-0
-
GD2-CAR T cell therapy for H3K27M-mutated diffuse midline gliomas
Majzner, R.G., Ramakrishna, S., Yeom, K.W. et al. GD2-CAR T cell therapy for H3K27M-mutated diffuse midline gliomas. Nature 603, 934–941 (2022). https://doi.org/10.1038/s41586-022-04489-4
-
Wang, H., Liu, C., Xie, X., Niu, M., Wang, Y., Cheng, X., Zhang, B., Zhang, D., Liu, M., Sun, R., Ma, Y., Ma, S., Wang, H., Zhu, G., Lu, Y., Huang, B., Su, P., Chen, X., Zhao, J., Wang, H., … Cheng, T. (2023). Multi-omics blood atlas reveals unique features of immune and platelet responses to SARS-CoV-2 Omicron breakthrough infection. Immunity, 56(6), 1410–1428.e8. https://doi.org/10.1016/j.immuni.2023.05.007

