Single-Cell Immune Repertoire Sequencing: Integrative Multi-Omics Dissection of the Immune Microenvironment
The 10× Genomics 5′ transcriptome combined with V(D)J sequencing enables the simultaneous acquisition of gene expression profiles and T/B cell receptor repertoires at single-cell resolution. When integrated with additional omics layers—including epigenomics, spatial transcriptomics, proteomics, and metabolomics—this approach allows for a systematic characterization of immune cell clonal dynamics, spatial organization, and regulatory networks. Such multi-dimensional profiling has emerged as a prevailing paradigm in high-impact research within the field of single-cell immunology1 (Figure 1).
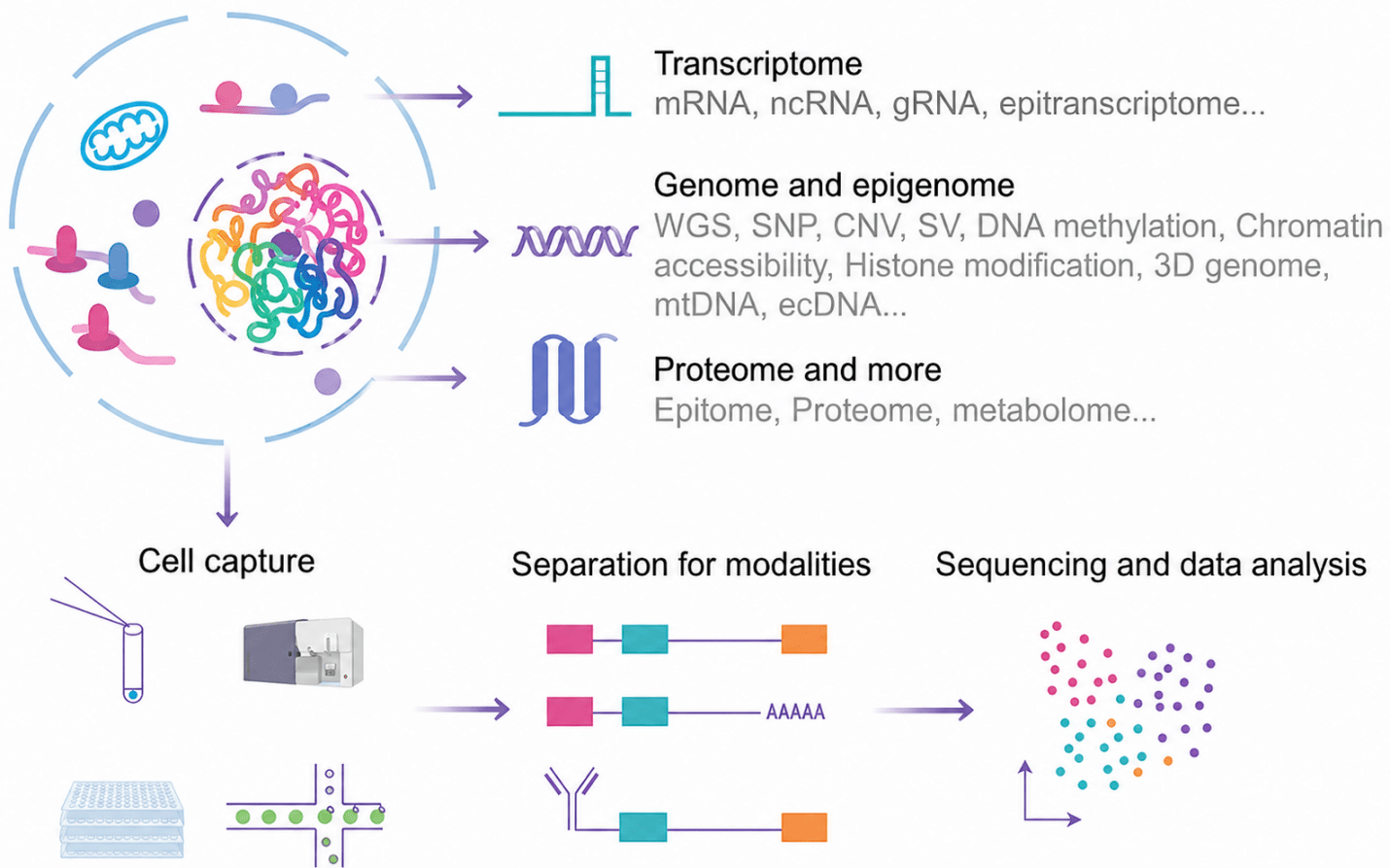
Figure 1. Schematic illustration of single-cell multi-omics
In contrast to conventional 3′ library construction strategies, the 10× Genomics 5′ transcriptome approach captures transcripts from the 5′ end of mRNA—thereby encompassing the V(D)J recombination regions of T and B cell receptors. This design enables the simultaneous acquisition of whole-transcriptome expression profiles and full-length immune receptor sequences within a single library preparation, without the need for additional amplification steps and without loss of receptor pairing information.
Furthermore, integrative multi-omics analysis—leveraging computational frameworks such as Seurat WNN, MOFA+, and ArchR—enables cross-modal cell embedding and reveals regulatory relationships that are not accessible through any single modality alone. This integrative strategy substantially enhances both the depth and innovation of biological discovery. Two representative case studies are presented below.
Case Study 1: Pan-Cancer Multi-Omics Profiling of Tumor-Associated Neutrophils
| Journal | Cell | Time | March 14, 2024 | Impact Factor | 42.5 |
| Multi-Omics Strategy | scRNA + V(D)J + Metabolomics + ATAC-Seq + CHIP-Seq + Spatial | ||||
Using 10× 5′ scRNA-seq as the backbone, the study integrated data from 17 cancer types and 225 samples to identify 10 transcriptional states of tumor-associated neutrophils, uncovering that the HLA-DR⁺ subset correlates with favorable prognosis across multiple cancers. But what drives this subset’s differentiation required other omics layers to answer.
Metabolomics + Epigenomics pinpointed the mechanism: untargeted metabolomics identified leucine metabolism as the key upstream driver; ATAC-seq and ChIP-seq then showed that leucine opens chromatin at MHC-II loci and upregulates H3K27ac modification—converting the metabolic signal into epigenetic reprogramming that shapes this neutrophil fate.
Spatial transcriptomics + V(D)J confirmed function in vivo: spatial data placed HLA-DR⁺ neutrophils directly within the tumor microenvironment, while V(D)J repertoire sequencing revealed that T cell clonal expansion co-localizes with these cells spatially—demonstrating that they drive genuine antigen-specific T cell responses, not bystander inflammation.
It is the single-cell resolution of 10× 5′ scRNA-seq that gives every downstream omics layer a shared anchor point—allowing metabolic, epigenetic, spatial, and immune repertoire findings to all converge on the same cell subset. Without it, the multi-omics relay has nowhere to land2.
Case Study 2: Pan-Cancer Multi-Omics Profiling of Tumor-Associated Neutrophils
| Journal | Immunity | Time | June 13, 2023 | Impact Factor | 26.3 |
| Multi-Omics Strategy | scRNA + V(D)J + Metabolomics + Proteomics + bulk RNA-seq | ||||
10× 5′ scRNA-seq coupled with BCR/TCR-seq served as the backbone of the study, enabling profiling of over 1,000 blood specimens across multiple disease phases, mapping the full cellular landscape of PBMC composition and capturing persistent B cell abnormalities that extended well into recovery. Yet the data kept pointing to an unexpected player—platelets.
Plasma proteomics + metabolomics took over: 447 differentially expressed metabolites and platelet-associated protein upregulation collectively established that platelets were not bystanders but active immune modulators—driving widespread platelet–leukocyte aggregate formation and amplifying interferon-mediated antiviral responses.

With the "what" identified, platelet bulk RNA-seq + BCR-seq dug into the "why" behind immune memory failure. Platelet transcriptomics confirmed sustained interferon dysregulation, while BCR repertoire data showed re-positive patients carried hallmarks of humoral exhaustion: collapsed clonality, diminished antibody output, and reduced Omicron neutralization. Folding all of this together with clinical indicators, the team trained a machine learning model capable of predicting re-positivity risk at the individual level.
Across every layer, the single-cell resolution of 10× 5′ scRNA-seq + BCR/TCR-seq served as the common thread—giving proteomics, metabolomics, and bulk transcriptomics a cellular context to anchor to, and transforming isolated molecular signals into a coherent map of the blood ecosystem3.
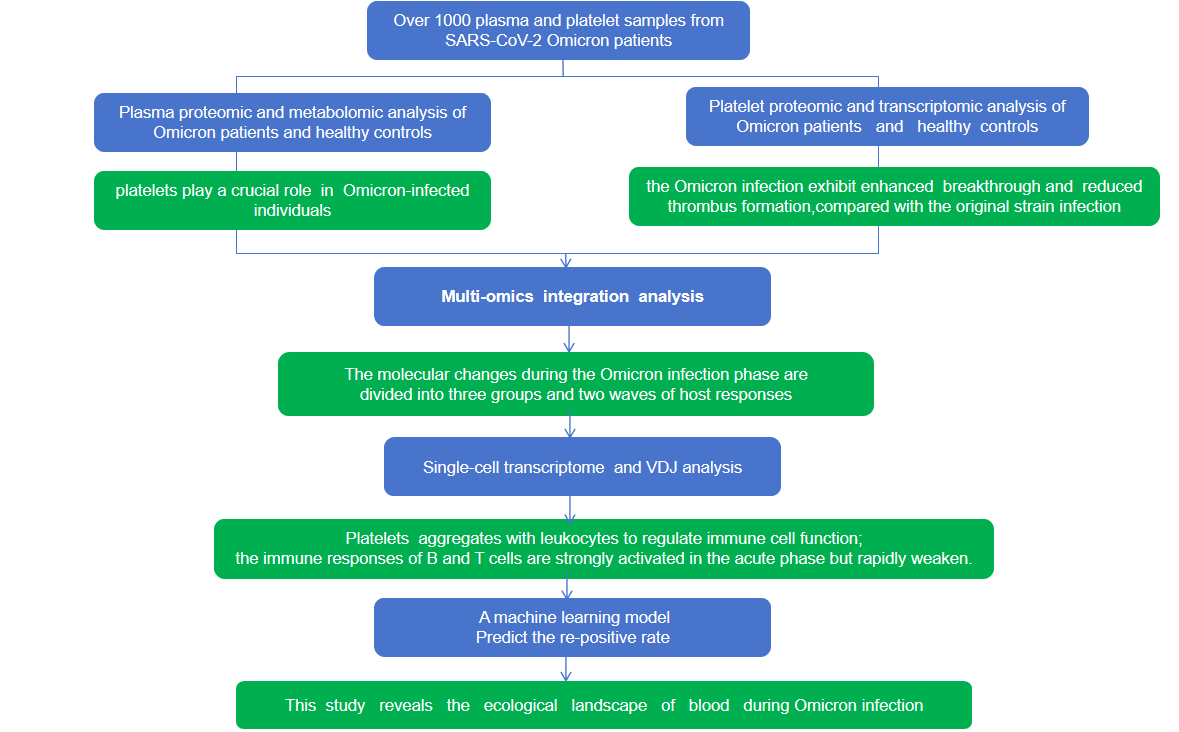
Figure 2. Case 2: Step-by-Step Research Framework
In summary, 10× 5′ transcriptomics combined with V(D)J sequencing offers a high-resolution, single-cell view into the complex regulatory networks of the immune system—making it an indispensable tool for both basic immunology research and clinical translation. When further integrated with proteomics, metabolomics, spatial transcriptomics, and epigenomics, the approach illuminates the underlying mechanisms of biological processes from multiple molecular dimensions simultaneously, accelerating the path toward precision medicine.
Reference
- Vandereyken K, Sifrim A, Thienpont B, Voet T. Methods and applications for single-cell and spatial multi-omics. Nat Rev Genet. 2023 Aug;24(8):494-515.
- WU Y, MA J, YANG X, et al. Neutrophil profiling illuminates anti-tumor antigen-presenting potency[J]. Cell, 2024, 187(6): 1422-1439.e24.
- WANG H, LIU C, XIE X, et al. Multi-omics blood atlas reveals unique features of immune and platelet responses to SARS-CoV-2 Omicron breakthrough infection[J]. Immunity, 2023, 56(6): 1234-1251.e8.
Why Choose Novogene for Single-cell RNA Sequencing (single-cell RNA-Seq or scRNA-Seq)?
Proven Expertise: With over 200,000 successfully sequenced samples, Novogene delivers great project results at industry-leading turnaround times. We excel at handling challenging sample types, including nerve and adipose cells.
Enhanced Sample Processing: We offer a diverse range of sample processing capabilities, including nuclei extraction and specialized pipelines for frozen tissues. This ensures high-quality gene expression data in Single-cell RNA Sequencing (single-cell RNA-Seq or scRNA-Seq) projects.
Certified Excellence: As a 10x Genomics Certified Service Provider, we leverage the advanced Chromium X platform combined with GEM-X technology for superior reproducibility and efficiency.
Cost-Effective Solutions: We have state-of-the-art high-throughput sequencing platforms, coupled with expert support, which ensure exceptional data quality and provide cost-effective solutions for single-cell projects.


