Metagenomics and Amplicon sequencing revealed soil biodiversity
Soil is not merely dirt; it has a complex biodiversity. Soil microbes dominate important processes, including plant growth and the cycle of elements such as carbon. Beneficial microorganisms in the soil carry out fundamental functions in nutrient cycling, breaking down crop residues, and stimulating plant growth. Soil conditions important for plant growth include the relative proportions of available nutrients (such as carbon [C], phosphorus [P], and Nitrogen [N]), minerals, and salinity. Furthermore, nematodes, bacteria, fungus, and viruses are only a few of the rich, intricate, and diverse multitrophic communities that may be found in soil. Plants have complex interactions with the biodiversity of their surroundings. Although the contribution of bacteria to crop production and soil health is clear, the biological component of the soil is very challenging to observe and manage.

Soil biodiversity and metagenomics
Different genomic applications have different strengths. Amplicon libraries are useful for quickly identifying advantageous mutations, analyzing genetic diversity in certain genomic regions, and assisting in selection decisions. Metagenomic sequencing technology is particularly useful for gathering information on the species and functional genes of microorganisms, and it captures microbial DNA from a variety of physiological states. Thus, metagenomics is valuable for investigating the soil microbes that dominate numerous crucial processes, such as plant growth and cycling of elements like C and P.
One important component of soil biodiversity is the rhizosphere, serving as a plant-root-soil microbiome interface. The rhizosphere is “the area around the roots that is directly influenced by root processes and that is the home of the rhizosphere microbiome.” (1) Thus, changes in the rhizosphere microbiome and bioavailability of nutrients in that space would be expected to have effects on plant growth and survival. In a 2021 study, Enebe and Babalola used shotgun metagenomic sequencing to investigate the effects of an organic P fertilizer source (compost) and/or inorganic fertilizer on P cycling genes in the maize rhizosphere microbiome. Their findings indicated that fertilization with organic compost and low amounts of inorganic fertilizer promoted microbial diversity involved in P cycling, whereas either high amounts of inorganic fertilizer or low amounts of compost fertilizer repressed microbial diversity involved in P cycling. (2) Ren et al. used metagenomic sequencing to investigate how soil microbes regulate C emissions in different forests. Their results suggested that expression profiles associated with labile C degradation drive soil priming, especially profiles associated with rapid simple sugar metabolism. (3)
The context and conditions of plant-rhizosphere microbiome relationships also matters. Increased incidence of drought is climate change-related concern, and therefore the resilience of plant-rhizosphere microbiome relationships should be investigated under those conditions. In a 2020 review, De Vries et al. proposed that to develop a more quantifiable, relevant understanding of plant-microbe interactions under drought conditions, future studies should focus on crop plants under real-world field conditions and use metagenomic analyses of controlled experiments. (1)
Studies like these emphasize the complexity of microbial diversity in the soil and of plant-microbe relationships. Further, they suggest that successfully increasing crop production in long-term, sustainable ways will require deeper understandings of the relationships between plants, soil microbes, soil conditions, and fertilizers. Yet there is much left to learn: many soil microorganisms cannot be isolated and cultured, and their functions and ecological roles remain unclear. Additionally, metagenomic analysis of gene function can only forecast a microbial community’s potential capabilities. The future of research will include meta phenotyping, which combines the genetic potential of the microbiome with its use of resources.
Effects of long-term anthropogenic activities on soil biodiversity
As discussed above, soil biodiversity encompasses multitrophic ecological networks that are affected by past and current farming practices, like long-term fertilizer use, and other anthropogenic activities. Advancing the goals of increased crop production also requires examining the effects of such anthropogenic activities on soil biodiversity and how that biodiversity impacts crop production.
In a 2020 study, Fan et al. used 16S rRNA and 18S rRNA amplicon sequencing to identify essential nematode, bacterial, and fungal phylotypes (keystone phylotypes) in wheat field soil following a 35-year fertilization experiment. Their results indicated that soils with more keystone phylotypes had higher wheat production, and that the keystone phylotypes expressed genes important for plants’ nutrient cycling. (16) Ma et al. used 16S rRNA amplicon sequencing to investigate the effects of reactive N deposition over 14 years on soil microbial communities. Their results suggested that N deposition resulted in significant changes in microbial communities above- and below ground. (4) These findings emphasize that strategies to increase future food production must consider and seek to better understand the impact of anthropogenic activities on soil biodiversity.
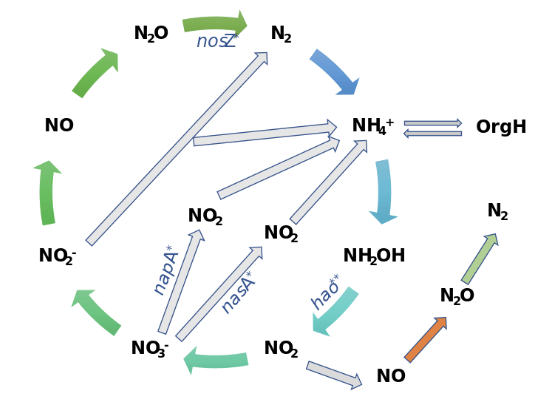
Changes in N cycling gene abundances. *P < 0.050; **P < 0.010. Modified from Ma X. et al. (2022) Long-term nitrogen deposition enhances microbial capacities in soil carbon stabilization but reduces network complexity. Microbiome 10:112.
Salt stress management
Strategies for improving crop growth in saline soil are important for two reasons. One, additional croplands may need to be developed to meet the need for increased agricultural production. (6) Two, climate change and rising temperatures cause increased soil salinity through increased evaporation of water from soil, which leaves salt behind in soil, as well as seawater intrusion in coastal zones. (6-7) Saline soil causes salt stress affects plant homeostasis through complex mechanisms that are not yet fully understood. (8) Strategies for improving growth in saline soil cover a variety of methods to which genomic approaches are applicable. Jiang et al. used 16S rRNA sequencing to test the effects of different fertilizer regimes on plant growth and soil microbiome composition. (6) Kong et al. used transcriptomics to investigate genome-wide differences in expression of salt stress-related genes in japonica (higher salt tolerance) and indica (lower salt tolerance) rice strains, finding salt stress-related candidate genes that may serve as gene editing targets to improve salt tolerance in rice. (8) Chauhan et al. also used transcriptomics, but to investigate the mechanisms driving beneficial rhizobacteria interactions with salt-stressed rice, suggesting B. amyloliquefaciens inoculation as a potential method for alleviating crops’ salt stress response. (7)
The soil biodiversity is closely related to the cycling of many elements such as soil organic matter, nitrogen and phosphorus, degradation of soil pollutants and inter-root immunity to soil-borne diseases. Genomic sequencing technologies, especially metagenomics and amplicon sequencing, enable efficient investigations of the complex relationships between crops, soil ecological networks, and their shared environments. For maintaining sustainable agricultural production and safeguarding the environment, it is crucial to advance research on the composition and function of the soil microbiome, strengthen its function in the soil ecosystem, achieve precise regulation of the soil microbiome and functional microbiome transplantation at specific or larger scales to reduce the application of chemical fertilizers and pesticides, and use microorganisms to remediate contaminated soils to improve soil health.
References
1. Vries F. T. D. et al. (2020) Harnessing rhizosphere microbiomes for drought‑resilient crop production. Science 368(6488):270.
2. Enebe M. and Babalola O. (2021) The influence of soil fertilization on the distribution and diversity of phosphorus cycling genes and microbes community of maize rhizosphere using shotgun metagenomics. Genes 12:1022.
3. Ren C. et al. (2021) Microbial traits determine soil C emission in response to fresh carbon inputs in forests across biomes. Global Change Biol. 28(4):1516.
4. Ma X. et al. (2022) Long‑term nitrogen deposition enhances microbial capacities in soil carbon stabilization but reduces network complexity. Microbiome 10:112.
5. Jiang S-Q. et al. (2019) High‑throughput absolute quantification sequencing reveals the effect of different fertilizer applications on bacterial community in a tomato cultivated coastal saline soil. Sci Total Environ. 687:601.
6. Chauhan P. S. et al. (2019) Transcriptional alterations reveal Bacillus amyloliquefaciens-rice cooperation under salt stress. Sci. Reports 9:11912.
7. Corwin D. L. (2020) Climate change impacts on soil salinity in agricultural areas. Eur. J. Soil Sci 72(2):842.
8. Kong W. et al. (2021) Comparative transcriptome analysis reveals the mechanisms underlying differences in salt tolerance between indica and japonica rice at the seedling stage. Front. Plant Sci. 12:725436.
