In the Lab: A Closer Look at DNA Methylation Sequencing Techniques
1 DNA Methylation: An Insights into the Epigenetic World
In the ever-evolving realm of molecular biology, the study of epigenetics has paved the way for groundbreaking discoveries in gene regulation and its profound impact on a wide array of biological processes. Among the most captivating facet of epigenetics lies in the DNA methylation. Imagine it as a maestro, influencing gene expression through a delicate dance of chromatin structure, DNA conformation, and protein interactions, and it is closely associated with physiological and pathological processes such as embryonic development [1], X chromosome inactivation [2], genome structural stability [3], genetic imprinting [4], aging [5], and the progression of tumors and other diseases [6].
Picture DNA methylation as a painter, dabbing colors onto the canvas of cytosine. DNA methylation can occur at various sites, including the C-5 position of cytosine, the N-4 position of cytosine, and the N-6 position of adenine, leading to the formation of C-5 position of cytosine, 5-hydroxymethylcytosine (5-hmC), N4-methylcytosine (N4-mC), and N6-methyladenine (N6-mA), and more. Among these, 5-mC takes the spotlight. It is often referred to as the "fifth base" of DNA and is widely present in the genomes of eukaryotic organisms. Meanwhile, 5-hmC is recognized as the "sixth base" of the mammalian genome [8] and plays a crucial role in the development of diseases, such as those affecting the nerve system and tumorigenesis [9, 10].
2 DNA Methylation: Sequencing Techniques
In the field of DNA methylation, a diverse array of methods, each with its unique strengths and limitations are available. The working principles of each of the sequencing techniques are quickly discussed below.
2.1 WGBS (Whole Genome Bisulfite Sequencing)
WGBS involves treating DNA with bisulfite, transforming unmethylated cytosines (C) to uracils (U). Subsequent PCR converts U to adenine (A), preserving only the methylated C. Following amplification, sequencing unveils genomic sites with methylation.
- Coverage: Offers a genome-wide methylation profile, allowing researchers to explore methylation patterns across all genomic regions.
- Applications: Valuable for fundamental research and clinical studies. Identifies epigenetic biomarkers and therapeutic targets for diseases.
- Cost: More expensive due to high sequencing depth requirements.
- Key Consideration: Researchers need to be aware of biases introduced during bisulfite conversion and PCR amplification.
2.2 RRBS (Reduced Representation Bisulfite Sequencing)
RRBS utilizes the restriction endonuclease Mspl enzyme to selectively cut the genome. After bisulfite treatment, sequencing in the specific regions such as gene promoters and CpG islands.
- Coverage: Does not provide a genome-wide view; selectively captures regions with high CpG density.
- Applications: Useful for targeted studies, such as identifying differentially methylated regions.
- Cost: Generally more cost-effective.
- Key Consideration: Choose RRBS when specific genomic regions are of interest.
2.3 MethylationEPIC v2.0 BeadChip (935K Chip)
The 935K Chip is a biochip adorned with millions of DNA probes, each matched to a CpG site in the genome. This chip acts like a molecular detective, spotlighting over 935,000 crucial CpG sites in regions like enhancers and promoters, revealing where DNA is methylated.
- Method: Microarray-based.
- Coverage: High-throughput profiling but may miss certain CpG sites.
- Applications: Enables large-scale screening of methylation patterns.
- Considerations: Limited to predefined CpG sites on the chip.
2.4 Enzymatic Methyl-sequencing (EM-seq)
EM-Seq transforms 5-mC and 5-hmC into 5-caC and 5-gmC using TET 2 and oxidation enhancers. APOBEC deaminase then converts unmethylated cytosines to Uracils (U). By studying these changes, methylated regions can be discerned.
- Method: Targeted approach capturing specific genomic regions with enzymatic precision.
- Applications: Provides focused insights into methylation patterns.
- Advantages: Precise and efficient.
2.5 Single-molecule real-time sequencing (SMRT-seq) and Nanopore-seq
SMRT-seq captures the essence of DNA sequencing by associating each base with a fluorescently labeled nucleotide during synthesis. This method provides a real-time picture of DNA synthesis, providing a dynamic understanding of the DNA sequence. Nanopore-seq utilizes nanoscale pores through which DNA passes, generating distinct electrical signals. As DNA molecules pass through the nanopores, different bases generate distinct electrical current signals, and then the sequencer determines the DNA base sequence.
- Method: New technologies providing long-read capabilities.
- Advantages: Long reads but may exhibit higher error rates.
- Considerations: Evaluate trade-offs between read length and accuracy.
A comparative overview of features of the DNA methylation technique is given in Table 1.
Table 1. Overview of different DNA methylation sequencing techniques
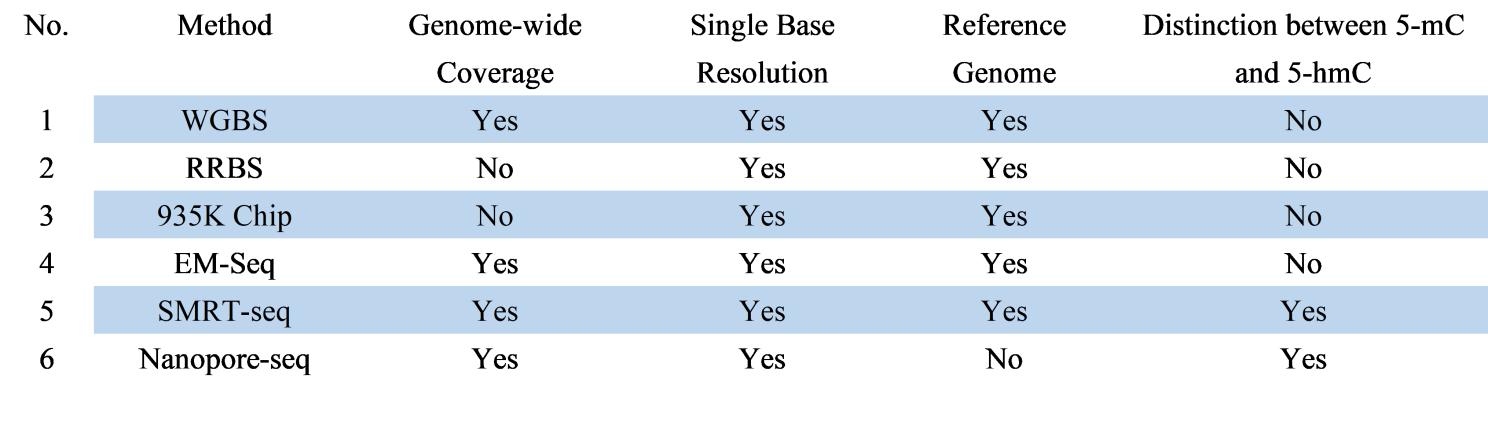
Remember that the choice of a DNA methylation sequencing method depends on your specific research objectives, available resources, and the balance between coverage, resolution, and accuracy. Each method has its strengths and limitations, so consider these factors carefully when designing your experiments.
3 Novogene’ s Whole Genome Bisulfite Sequencing (WGBS) Services
Novogene’s Whole Genome Bisulfite Sequencing (WGBS) seamlessly integrates the bisulfite treatment method with a high-throughput sequencing platform, enabling comprehensive methylation studies at the whole-genome level for species with a reference genome. Recent data from Novogene underscores exceptional efficiency, boasting a bisulfite conversion rate of 99.65% and a library construction success rate surpassing 99.66%. These impressive figures ensure the generation of high-quality research outcomes. Novogene boasts extensive experience in methylation sequencing analysis across a diverse array of species, from pigs, rhesus macaques, and Yangtze alligators to oysters, abalones, Arabidopsis, rice, chili peppers, poplar trees, and beyond. This rich background has empowered researchers partnered with Novogene to publish numerous high-quality articles in authoritative journals.
Novogene provides an impressive array of comprehensive services, ensuring top-notch quality and cutting-edge analysis. Services encompass the following:
- Sequencing data quality control
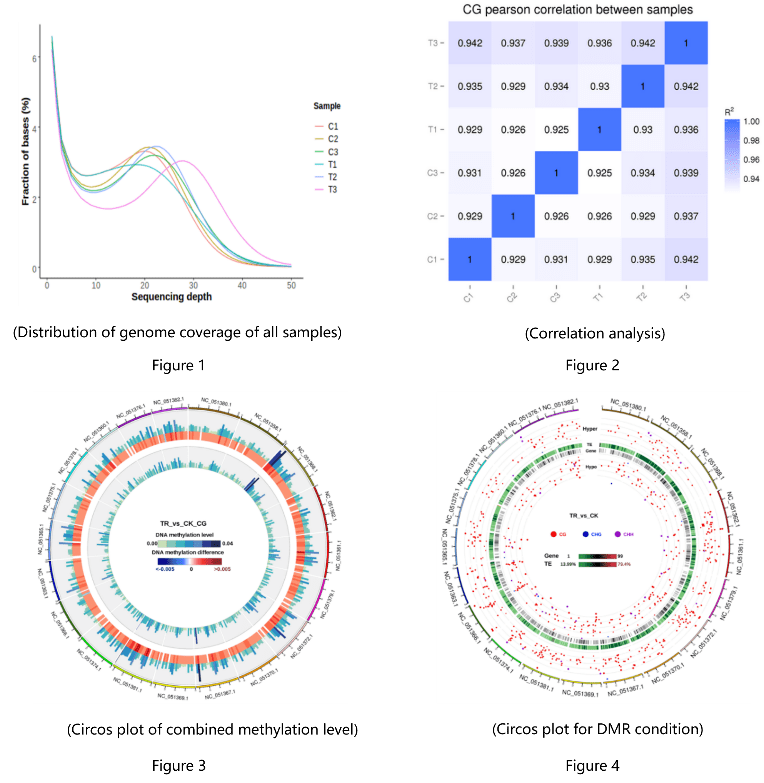
- Reference sequence alignment analysis (Figure 1)
- Sample correlation analysis (Figure 2)
- Genome-wide methylation site detection and analysis (Figure 3)
- Evaluation of methylated C sites in various sequence contexts (CG, CHH, CHG)
- Functional region examination
- Differential methylation region (DMR) analysis (Figure 4)
- Distribution of DMR-anchored regions
- GO enrichment analysis and KEGG enrichment analysis of DMR-related genes (Figure 5-6)
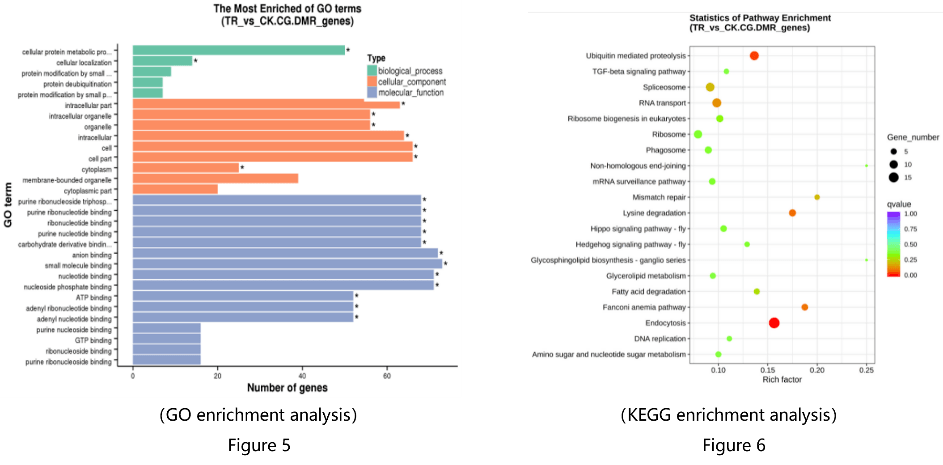
As a trusted research partner, Novogene is wholeheartedly committed to providing exceptional support. With professional pre-sales and after-sales technical teams and an after-sales cloud platform (Customer Service System, CSS), Novogene promptly addresses various research needs and personalized analysis issues. For any specific needs, researchers are encouraged to reach out to Novogene for prompt assistance.
4 Reference
[1] WU X, ZHANG H, ZHANG B, et al. Methylome inheritance and enhancer dememorization reset an epigenetic gate safeguarding embryonic programs [J]. Science advances, 2021, 7(52): eabl3858.
[2] CANTONE I, FISHER A G. Human X chromosome inactivation and reactivation: implications for cell reprogramming and disease [J]. Philosophical transactions of the Royal Society of London Series B, Biological sciences, 2017, 372(1733): 20160358.
[3] HE L, ZHAO C, ZHANG Q, et al. Pathway conversion enables a double-lock mechanism to maintain DNA methylation and genome stability [J]. Proceedings of the National Academy of Sciences of the United States of America, 2021, 118(35).
[4] GREESON K, CROW K, EDENFIELD R, et al. Inheritance of paternal lifestyles and exposures through sperm DNA methylation [J]. Nature reviews Urology, 2023, 20(6): 356-70.
[5] SEALE K, HORVATH S, TESCHENDORFF A, et al. Making sense of the ageing methylome [J]. Nature reviews Genetics, 2022, 23(10): 585-605.
[6] ROY D, TIIRIKAINEN M. Diagnostic Power of DNA Methylation Classifiers for Early Detection of Cancer [J]. Trends in cancer, 2020, (2): 6.
[7] Advances in mapping the epigenetic modifications of 5-methylcytosine (5mC), N6-methyladenine (6mA), and N4-methylcytosine (4mC) [J]. Biotechnology and Bioengineering.
[8] MüNZEL M, GLOBISCH D, CARELL T. 5‐Hydroxymethylcytosine, the Sixth Base of the Genome [J]. Angewandte Chemie International Edition, 2011, 50(29).
[9] SZULWACH K E, LI X, LI Y, et al. 5-hmC-mediated epigenetic dynamics during postnatal neurodevelopment and aging [J]. Nature Neuroscience, 2011, 14(12): 1607-16.
[10] EHRLICH M, LACEY M. DNA Hypomethylation and Hemimethylation in Cancer [J]. Advances in Experimental Medicine and Biology, 2013, 754: 31-56.
