Exploring the Power of Whole Genome Sequencing: A Beginner’s Guide to Long-read sequencing
Exploring the Power of Whole Genome Sequencing: A Beginner’s Guide to Long-read sequencing
In recent years, whole genome sequencing (WGS) has revolutionized the field of genetics and opened up new avenues for scientific exploration. It allows researchers to unravel the mysteries encoded within the entire genetic makeup of an organism, be it human, plant, animal, or microbe. WGS involves deciphering the full spectrum of genetic variation present within an organism’s entire genome. However, commonly used short-read sequencing (SRS) techniques for WGS have limitations in accuracy, read length, and interpreting repetitive sequencing. Long-read sequencing (LRS), a recent breakthrough technique, gives new hope for resolving these hurdles.
- Next-Generation Sequencing (NGS)
NGS, particularly represented by Illumina SBS sequencing technology, offer short-read sequencing technology, which is widely used for WGS due to its high-throughput capabilities and cost-effectiveness. It involves the parallel sequencing of short DNA fragments, generating millions of reads that are subsequently assembled to reconstruct the entire genome.
- Third-Generation Sequencing (TGS)
TGS approaches, such as Nanopore PromethIon and PacBio sequencing technologies, offer long-read sequencing technology, which enables the sequencing of much longer fragments of DNA than is possible with NGS. TGS methods are particularly useful for capturing information about complex genomic regions, resolving structural variations, and improving genome assembly
Why we would benefit to use WGS with third-generation sequencing
Long-read sequencing has substantial benefits over earlier short-generation sequencing approaches and is poised to transform WGS. It delivers whole genome sequence information with complete and accurate resolution through unbiased, reads length of up to 25 kb, a median read accuracy of Q30 (>99.9%), and the ability to sequence through repeats and GC-rich regions. By significantly reducing error rates, ensuring reliable results of longer read length and precise interpretation, it enables a better understanding of complex genomic structures such as gene rearrangements and transposons. These advancements will pave the way for further breakthroughs in genomics, driving progress in the life sciences, medical research, and personalized healthcare [1].
Fig 1. Comparison and specification of sequencing platforms between short-read and long-read sequencing on WGS
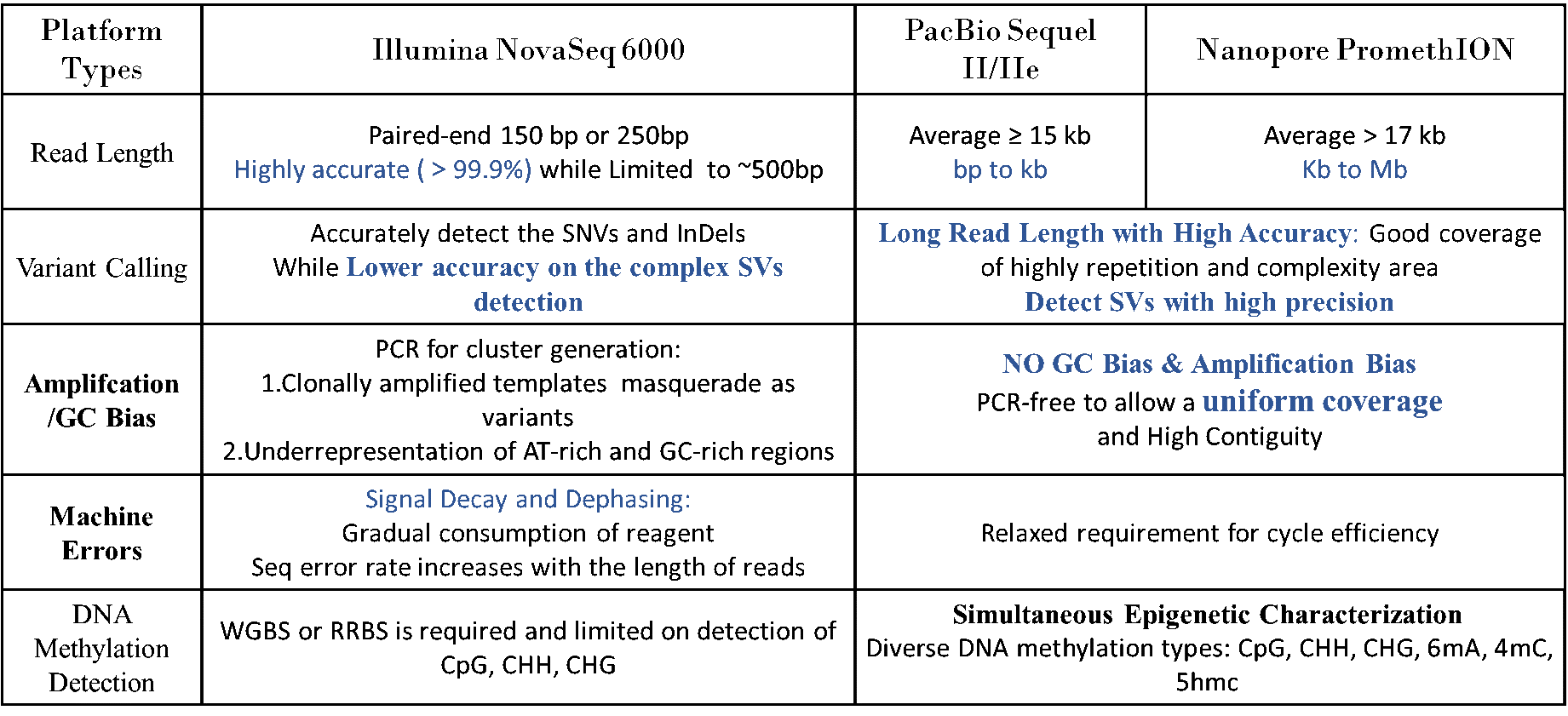
What kind of biological questions of interest can be addressed by WGS using Long-read sequencing?
Whole genome sequencing (WGS) using long-read sequencing has the potential to address a wide range of biological questions.
- Structural variations and genome rearrangements: Long-read sequencing accurately identifies and characterizes large-scale structural variations, such as deletions, duplications, inversions, and translocations. This enables researchers to understand their impact on gene function, disease susceptibility, and evolutionary processes.[2]
- Repeat expansion disorders: Long-read sequencing accurately captures repetitive DNA sequences, allowing for precise identification and sizing of repeat expansions. This is critical in unraveling the molecular basis of repeat expansion disorders, including Huntington’s disease and fragile X syndrome.
- Haplotype phasing and genome assembly: Long-read sequencing generates longer reads that span multiple genetic variants, facilitating accurate haplotype phasing and improving genome assembly quality. This information enhances our understanding of complex genetic traits, disease-causing variants, and genome structure.
- Comparative genomics and evolutionary studies: Long-read sequencing facilitates detailed genome comparisons across species or populations, unveiling insights into genomic rearrangements, variation patterns, and evolutionary relationships. This contributes to our understanding of species diversification, adaptation, and the identification of genes under positive selection.
By overcoming the limitations of short-read sequencing, long-read technology enables a comprehensive exploration of the genome’s intricacies and the underlying mechanisms driving biological phenomena.
A case study of neurological disorders investigation applying long-read sequencing supported by Novogene
The study was to explore whether a particular gene repeat expansion mutation is associated with the neurological disorder known as essential tremor (ET) [3]. This disorder has a strong genetic component, with more than half of cases reporting a family history of the condition. However, the identification of pathogenic genetic variants associated with sporadic cases of ET has been challenging, and the underlying pathogenic mechanism remains elusive.
In this study, LRS was employed to investigate whether GC repeat expansions of the NOTCH2NLC gene are associated with sporadic cases of ET. An additional objective was to determine whether these repeat expansions could be identified in clinically well-characterized familial ET pedigrees.
ET cases were screened for NOTCH2NLC GGC repeat expansions using repeat-primed polymerase chain reaction (RP-PCR). Cases that showed positive or intermediate repeat expansions were sent for confirmation using long-read sequencing was used, which is capable of detecting single nucleotide variants (SNVs), in complex regions of the genome, with high sensitivity and specificity. This is not possible with NGS, due to the short sequencing length used. Next, the study team performed primer fluorescence amplification refractory mutation system (ARMS) amplicon long-read analysis to characterize structural variants of 50 or more base pairs. The characterization of structural variants is crucial in genetic studies of neurological disease; however, the short-length sequencing used in NGS has again previously imposed a severe limitation on the study of these complex structural variants, particularly those involving repetitive regions. Therefore, LRS is essential to study the occurrence of large structural variants. Without the use of long-read sequencing, it would not have been possible to perform the fine-grained analysis of the NOTCH2NLC gene used in this study and determine the association of GC repeat expansions in this gene with sporadic cases of ET.
Long-read sequencing technology is propelling the investigation of neurological disorders to a new height. Novogene, as a renowned provider of genomics services, is proud to enable this research by leveraging state-of-the-art sequencing technologies, advanced bioinformatics analysis capabilities, the extensive knowledge and experience, and the trusted services. Novogene is committed to provide their clients with innovative solutions and valuable insights. These insights are helping to transform our understanding of previously inscrutable genetic disorders.
References:
1. Chen, X., et al., Whole-genome resequencing using next-generation and Nanopore sequencing for molecular characterization of T-DNA integration in transgenic poplar 741. BMC Genomics, 2021. 22(1): p. 329.
2. Functional Genomics Center Zurich. Whole Genome Resequencing. nd; Available from: https://fgcz.ch/omics_areas/genomics_uc/applications/whole-genome-resequencing.html.
3. Ng, A.S.L., et al., NOTCH2NLC GGC Repeat Expansions Are Associated with Sporadic Essential Tremor: Variable Disease Expressivity on Long-Term Follow-up. Ann Neurol, 2020. 88(3): p. 614-618.
